Mountains Of Data Being Used To Combat Citrus Greening
Across the globe, researchers are conducting hundreds of scientific studies aimed at finding solutions to HLB. The amount of data this work is generating is staggering, creating a challenge in and of itself. How can researchers seeking the latest knowledge on the disease find it in a sea of information that is growing by leaps and bounds on a daily basis?
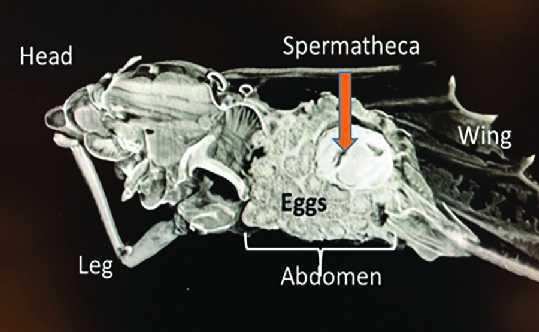
Literally, millions of CT Scan images are being generated of the Asian citrus psyllid as scientists study the pests. Photo courtesy of USDA
“Researchers today produce more data in a few months than science has produced in all the years previously,” says Wayne Hunter, a research entomologist at USDA’s Horticultural Lab in Ft. Pierce. “Today’s science depends heavily on computers, and in modern biology, every week is big data week. Lifesciences research now routinely churns out more information than scientists can analyze without the help of specialists and/or special software on high-speed computers.
“So while a lot of data is produced, and there is a lot of HLB-associated information, if we cannot access it, or analyze it efficiently, we are not using our full potential to solve the HLB problem.”
Getting A Handle On Analytics
Hunter says to conceptualize how researchers will find better ways to manage data in fields like ag science requires a shift from traditional ways of searching in a linear path to using more of a circular path or a spider web of connected information. The system would store data and provide common and secure access methods that would allow all those working on a problem from around the globe to query information.
“The data system must have an easy way to query or search for information, which permits using basic words like ‘bug’ or ‘disease,’ as well as more technical search terms like ‘Diaphorina’ or ‘Liberibacter,’ and then retrieve a range of information designed to be used by the layperson, grower, educator, politician, or researcher.
“Emerging digital databases also must be able to cope with data in different locations — on remote servers, on desktops, in a database, or spread across different machines — and in a wide range of formats, including spreadsheets, badly named files, blogs, or even scanned-in notebooks. And, of course, it will be helpful if the information can be accurately translated into other languages, like Spanish, or from other languages into English. There is still a lot of information on HLB, but it’s in Chinese, so it’s unavailable to most U.S. researchers.”
Building A System
Currently, Hunter is working with a team to build an open source portal on the Internet to be a clearinghouse of information on the trials attempting to culture the HLB bacteria, C. Liberibacter asiaticus (CLas). The bacteria has proven extremely difficult to culture in the lab, which would be a major breakthrough toward finding solutions to HLB.
“While many labs have tried, only two have proposed and published any tentative success,” he says. “To really understand how to culture this bacterium, it would be immensely helpful if all the trials and all their methods and results had been published in one location. That way, others can see what was done and how it was done. Then new experiments could be designed that would have a higher chance of success. To these ends, we have formed a team to start this process.”
The team also is preparing a publication on using Micro CT Scanning technology to provide 3D digital representations of the Asian citrus psyllid’s anatomy. Many species descriptions, especially older ones, consist mostly of text and have few illustrations. The publication will propose that descriptions should become more data-rich by presenting a large amount of images and illustrations to cover as much morphology as possible.
“We also have submitted for a National Institute Of Food And Agriculture grant, which incorporates many disciplines on HLB from genomics, proteomics, RNAi, molecule delivery, citrus breeding, and education to obtain funding for establishing ‘A Systems Biology Digital Database’ for the benefit of the citrus industry,” Hunter says. “The database will bring together — and most importantly — link the data together by associated clustering algorithms for a new and more rapid approach in examining and understanding the associations between various types of research and grower information. It will link the anatomical data, the genomic data (genes and proteins), pathogen movement, transmission, and survival. It also will start a local repository in Florida for all information associated with the psyllid, HLB, and citrus, which will be more user friendly for education, public, political, or researcher needs.”
While this system will be dedicated to helping solve the citrus greening problem, the database is planned to support all aspects of citrus research and education future needs and will be adaptable for current and emerging pests, disease pathogens, and citrus management practices. This system could serve as the foundation for managing and searching for data associated with other agricultural science research in other problems and crops.










